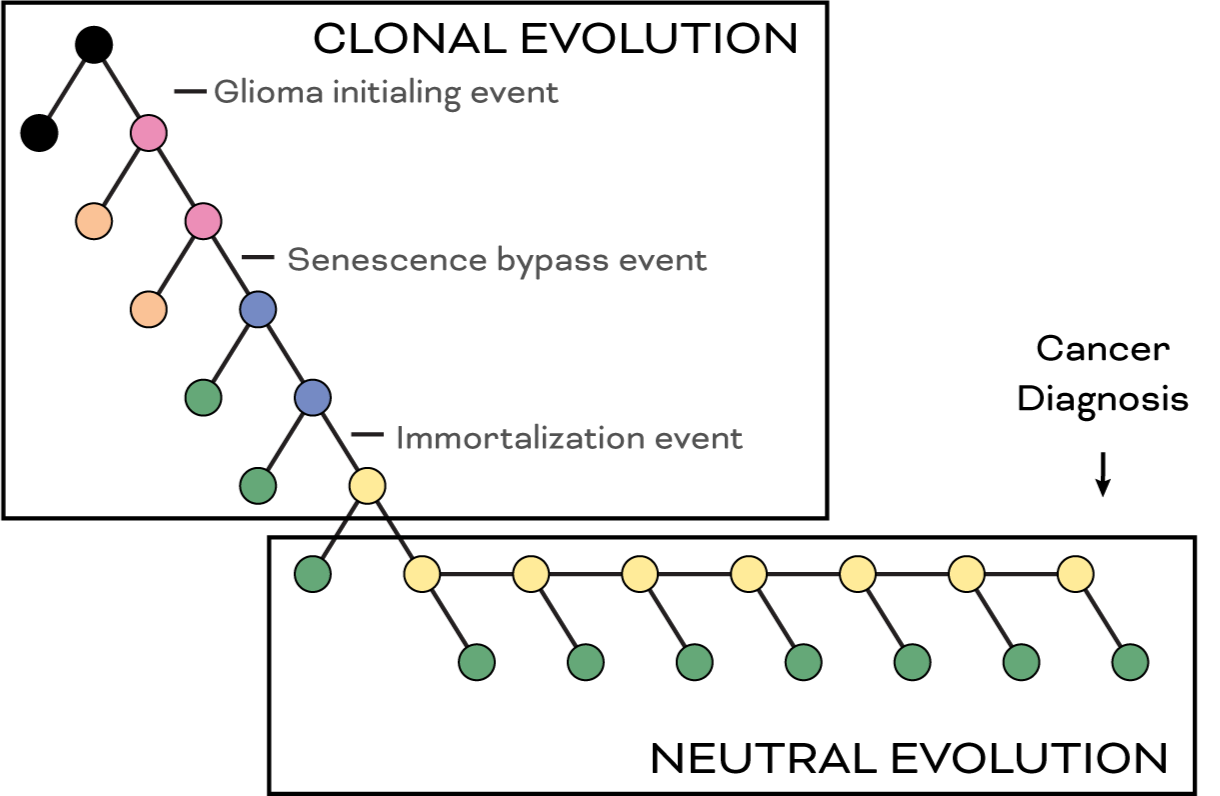
The Barthel laboratory studies glioma development and evolution. We employ computational biology, evolutionary genetics, functional genomics and molecular biology to study tumor development, treatment effect and tumor progression. We are looking for a talented, passionate and highly motivated individual to enrich our team and work on tumor evolution and longitudinal tracking of disease from liquid biopsies.
Position Summary
This project aims to reconstruct evolutionary and developmental lineages of gliomagenesis, as well as track disease progression by analyzing liquid biopsies from blood and cerebrospinal fluid (CSF). The successful applicant will employ next-generation genomic analysis (deep DNA sequencing, epigenome profiling, single-cell RNA sequencing) across multiple image-mapped samples per tumor to reconstruct tumor phylogenies and evolutionary trajectories in space and time. Serum/plasma and CSF samples will be deeply characterized to identify biomarkers of disease progression and treatment effect. They will collaborate with other scientists in the lab to detect telomere dysfunction in patient samples and model systems and study the evolution of telomere dysfunction driven structural variation. They will be involved in data analysis, interpretation and write manuscripts to describe their findings. They will lead their own project but will be a part of a team and help on other projects as well.
Career Development
We value our trainee’s personal development and strive to unlock their full potential. Trainees are provided with ample opportunities for additional training in computational and cancer biology. Successful applicants are encouraged to attend scientific meetings and the lab will provide support for at least one scientific conference per year. While this position is not contingent upon securing additional funding, successful candidates will be supported and encouraged to apply for outside sources of funding. Candidates are welcomed to explore their own scientific interests into areas of research that fall within the general scope of the lab.
Education and Preferred Qualifications
The ideal candidate will have a PhD in a relevant field and have experienced analyzing next-generation sequencing data. A track record of peer reviewed publications in (tumor) evolution is preferred. An understanding of cancer biology and genomics is a plus. The candidate should have experience with a statistical programming language such as Python or R and have a good understanding of statistical theory, phylogenetics and evolution. Experience in Unix-based computing environments is a plus. Candidates from all backgrounds will be considered.
Location
We are located at the Translational Genomics Institute (TGen) set to the backdrop of the stunning Sonoran Desert. The successful candidate will be a part of the Cancer and Cell Biology Division at TGen and will have numerous opportunities to work with our partners at around the globe.
Application
Please direct all questions to barthel@barthel-lab.com. Application via the “Apply” button on the left menu. Attach a letter describing your research interests and motivation to join our group. Include an academic CV detailing your past research experience, publication track record and (conference) presentations. We will start reviewing applications immediately and until the position is filled.